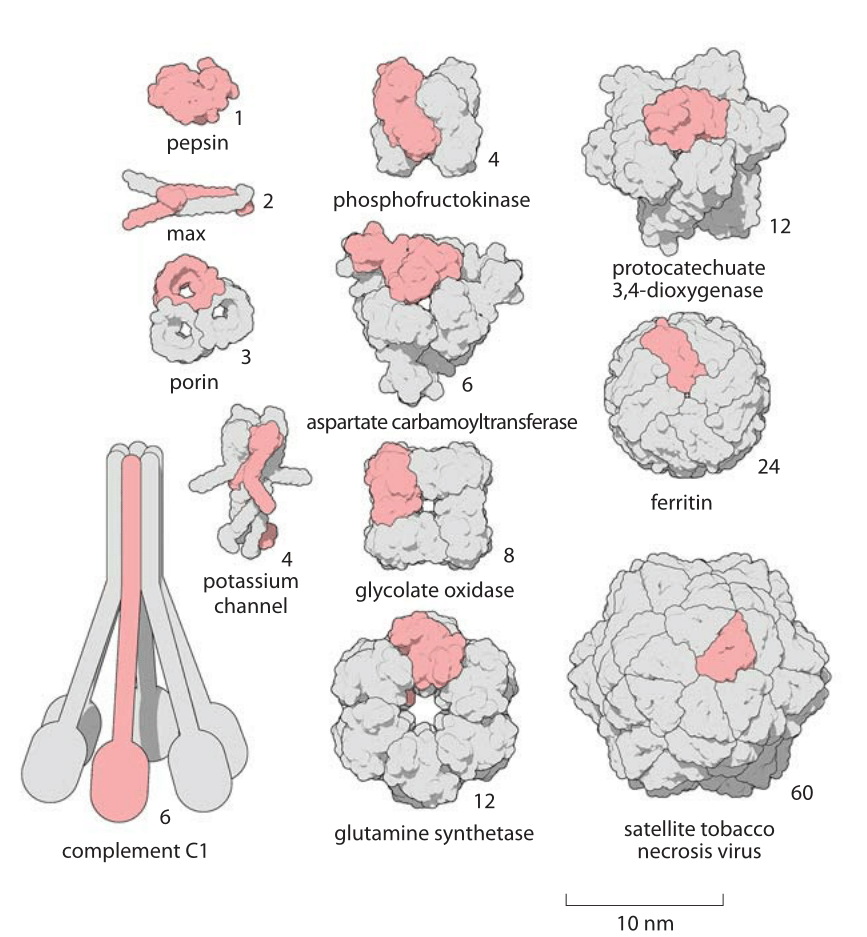
Running protein structure prediction at scale using a web interface for researchers
$ 41.00
-
By A Mystery Man Writer
-
-
4.6(541)

Product Description

Protein domain - Wikipedia

NAMD - Scalable Molecular Dynamics

Improved protein structure prediction using predicted interresidue

Running protein structure prediction at scale using a web

AI-Based Protein Structure Prediction in Drug Discovery: Impacts

ColabFold - Run Protein Prediction Locally

Evaluating AlphaFold protein-protein binding with ChimeraX
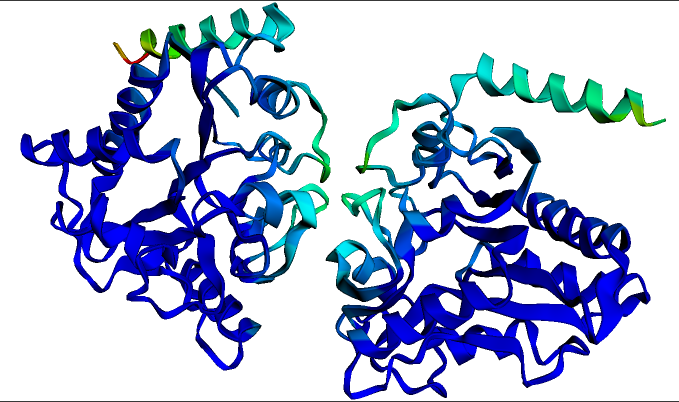
Deep Learning with Proteins
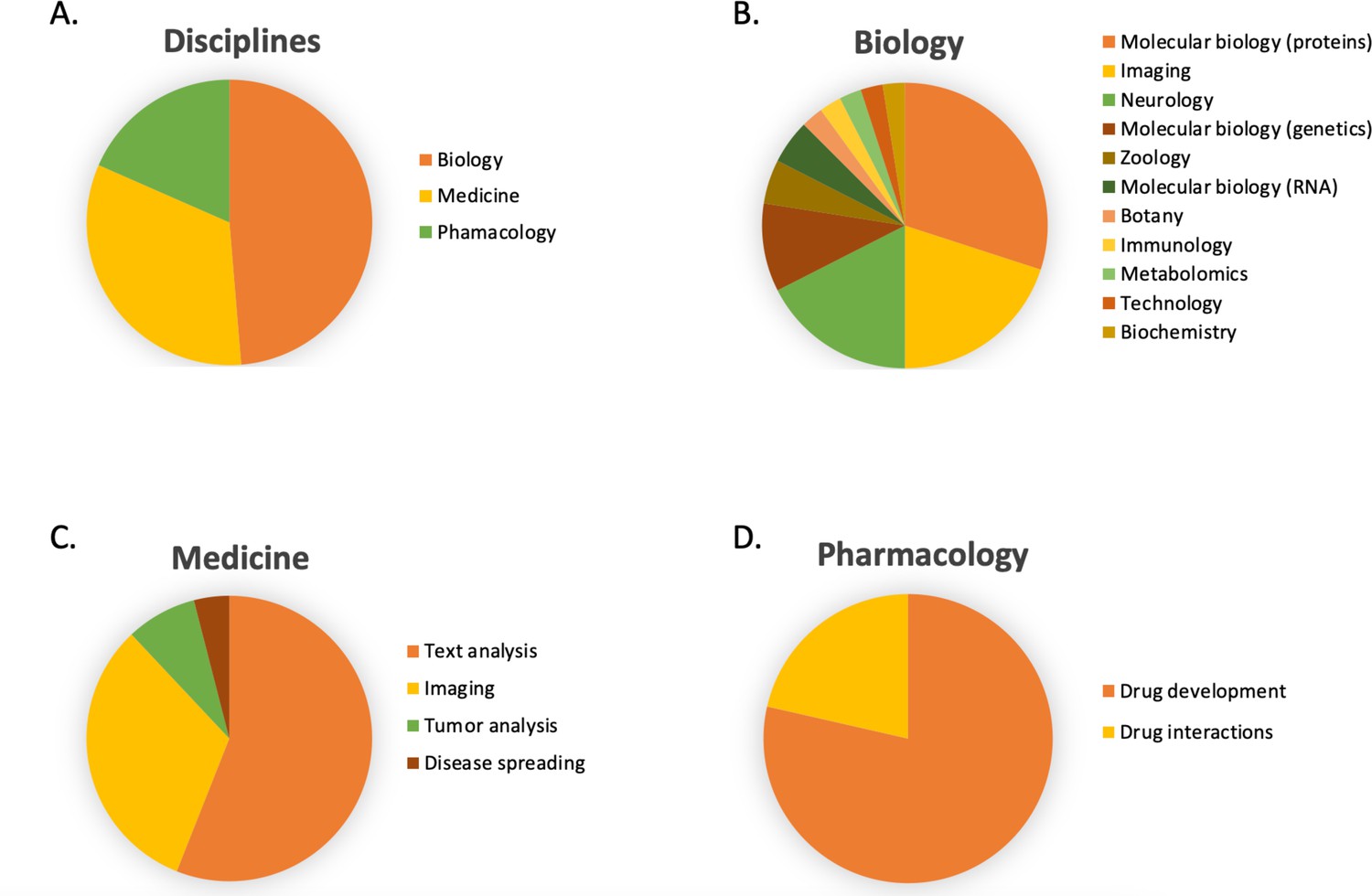
Transformer-based deep learning for predicting protein properties
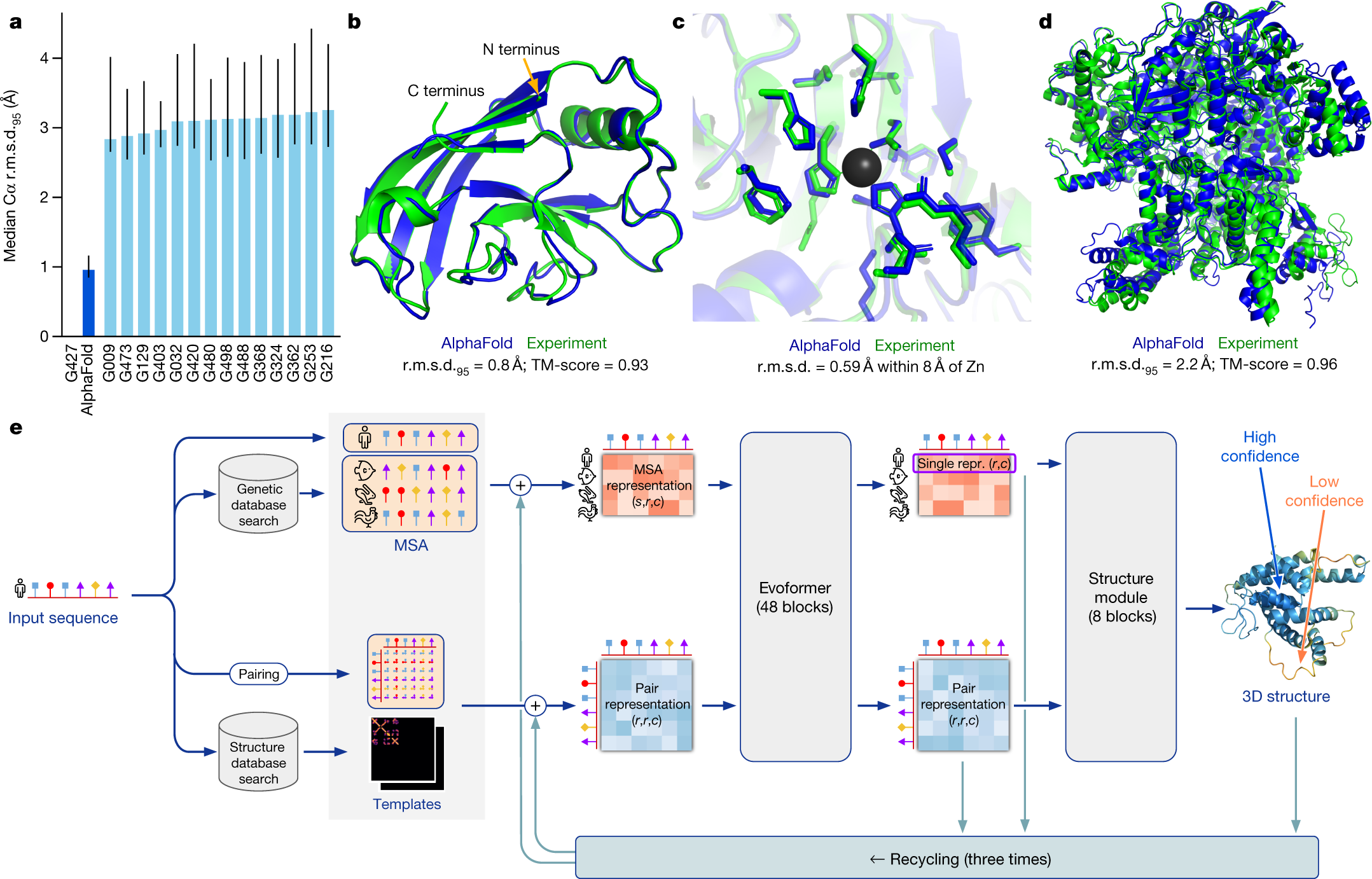
Highly accurate protein structure prediction with AlphaFold

AlphaFold, Artificial Intelligence (AI), and Allostery